Paolo Cascio
Professore/Professoressa ordinario/a
- Dipartimento di Scienze Veterinarie
- SSD: BIO/10 - biochimica

Contatti
- +39 0116709109
- paolo.cascio@unito.it
- Dipartimento di Scienze Veterinarie
Largo Paolo Braccini 2
10095, Grugliasco (TO), Italia - https://people.unito.it/persone/paolo.cascio
- VCard contatti
- QRcode contatti
Presso
- Department of Veterinary Sciences
- Dipartimento di Scienze Veterinarie - DSV
- Corso di Laurea in Produzioni e Gestione degli Animali in Allevamento e Selvatici - PGAAS
- Corso di Laurea in Sistemi zootecnici sostenibili - SIZOS
- Corso di Laurea in Tecniche di assistenza veterinaria - TAVET
- Degree Course in Veterinary Medicine
- Laurea Magistrale a Ciclo Unico in Medicina Veterinaria
Curriculum vitae
 Curriculum Vitae
(0.3 MB)
Curriculum Vitae
(0.3 MB) Prodotti della ricerca selezionati
Peer-reviewed publications
- Cerruti F., Borrelli A., Degiovanni A., Mengozzi G., Borella F., Cascio P. Detection and biochemical characterization of circulating proteasomes in dog plasma. Research in Veterinary Science, 2023, 162, 104950 (I.F. 2022 2.4)
- Tundo G.R., Cascio P., Milardi D., Santoro A.M., Graziani G., Lacal P.M., Bocedi A., Oddone F., Parravano M., Coletta A., Coletta M., Sbardella D. Targeting immunoproteasome in neurodegeneration: A glance to the future. Pharmacology & Therapeutics, 2023, 241, 108329 (I.F. 2022 13.5)
- Sbardella D., Tundo G.R., Mecchia A., Palumbo C., Atzori M.G., Levati L., Boccaccini A., Caccuri A.M., Cascio P., Lacal P.M., Graziani G., Varano M., Coletta M., Parravano M. A novel and atypical NF-KB pro-inflammatory program regulated by a CamKII-proteasome axis is involved in the early activation of Muller glia by high glucose. Cell & Bioscience, 2022, 12:108 (I.F. 7.5)
- Cantiello M., Carletti M., Giantin M., Gardini G., Capolongo F., Cascio P., Pauletto M., Girolami F., Dacasto M., Nebbia C. Induction by Phenobarbital of Phase I and II Xenobiotic-Metabolizing Enzymes in Bovine Liver: An Overall Catalytic and Immunochemical Characterization. International Journal of Molecular Sciences, 2022, 23, 3564 (I.F. 5.6)
- Cascio P. and Dittmar G. Regulating Proteasome Activity. Biomolecules, 2022, 12, 343 (I.F. 5.5)
- Tundo G.R., Sbardella D., Oddone F., Grasso G., Marini S., Atzori M.G., Santoro A.M., Milardi D., Bellia F., Macari G., Graziani G., Polticelli F., Cascio P., Parravano M., Coletta M. Insulin-Degrading Enzyme Is a Non Proteasomal Target of Carfilzomib and Affects the 20S Proteasome Inhibition by the Drug. Biomolecules, 2022, 12, 315 (I.F. 5.5)
- Frayssinhes J-Y.A., Cerruti F., Laulin J., Cattaneo A., Bachi A., Apcher S., Coux O., Cascio P. PA28γ-20S proteasome is a proteolytic complex committed to degrade unfolded proteins. Cellular and Molecular Life Sciences, Published online 2021 Dec 16;79(1):45 (I.F. 9.2)
- Cascio P. PA28γ: new insights on an ancient proteasome activator. Biomolecules, 2021, 11, 228 (I.F. 6.1)
- Boulpicante M., Darrigrand R., Pierson A., Salgues V., Rouillon M., Gaudineau B., Khaled M., Cattaneo A., Bachi A., Cascio P., Apcher S. Tumors escape immunosurveillance by overexpressing the proteasome activator PSME3. OncoImmunology, 2020, 9(1): 1-16, 1761205. (I.F. 8.1)
- Bordini J., Morisi F., Cerruti F., Cascio P., Camaschella C., Ghia P., Campanella A. Iron causes lipid oxidation and inhibits proteasome function in multiple myeloma plasma cells: a proof of concept for novel combination therapies. Cancers, 2020, 12(4): 970. (I.F. 6.6)
- Cerruti, F., Jocollè, G., Salio, C., Oliva, L., Paglietti, L., Alessandria, B., Mioletti, S., Donati, G., Numico, G., Cenci, S., Cascio, P. Proteasome stress sensitizes malignant pleural mesothelioma cells to bortezomib-induced apoptosis. Scientific Reports, 2017, 7: 17626. (I.F. 4.1)
- Oliva L., Orfanelli U., Resnati M., Raimondi A., Orsi A., Milan E., Palladini G., Milani P., Cerruti F., Cascio P., Casarini S., Rognoni P., Touvier T., Marcatti M., Ciceri F., Mangiacavalli S., Corso A., Merlini G., Cenci S. The amyloidogenic light chain is a stressor that sensitizes plasma cells to proteasome inhibitor toxicity. Blood, 2017, 129(15): 2132-42. (I.F. 15.1)
- Crespo H., Bertolotti L., Proffiti M., Cascio P., Cerruti F., Acutis P.L., de Andrés D., Reina R., Rosati S. Low proviral small ruminant lentivirus load as biomarker of natural restriction in goats. Microbiology, 2016, 192: 152-62. (I.F. 2,6)
- Milan E., Perini T., Resnati M., Orfanelli U., Oliva L., Raimondi A., Cascio P., Bachi A., Marcatti M., Ciceri F., Cenci S. A plastic SQSTM1/p62-dependent autophagic reserve maintains proteostasis and determines proteasome inhibitor susceptibility in multiple myeloma cells. Autophagy, 2015, 11(7): 1161-78. (I.F. 9,1)
- Raule M., Cerruti F., Cascio P. Comparative study of the biochemical properties of proteasomes in domestic animals. Veterinary Immunology and Immunopathology, 2015, 166(1-2): 43-9. (I.F. 1,7)
- Cascio P., Cerruti F., Marshall R.S., Raule M., Remelli W., Roberts L.M., Ceriotti A. A Quantitative Method to Monitor the Efficacy of Inhibitors Against the Chymotrypsin-Like Activity of the Proteasome in Tobacco Leaf Protoplasts. Plant Molecular Biology Reporter, 2015, 33(4): 829-840 (I.F. 2,3)
- Cascio P. PA28αβ: The Enigmatic Magic Ring of the Proteasome? Biomolecules, 2014, 4: 566-584. (I.F. N.A.)
- Raule M., Cerruti F., Cascio P. Enhanced rate of degradation of basic proteins by 26S immunoproteasomes. Biochimica et Biophysica Acta - Molecular Cell Research, 2014, 1843: 1942-47. (I.F. 5,0)
- Raule M., Cerruti F., Benaroudj N., Migotti R., Kikuchi J., Bachi A., Navon A., Dittmar G., Cascio P. PA28αβ Reduces Size and Increases Hydrophilicity of 20S Immunoproteasome Peptide Products. Chemistry & Biology, 2014, 21: 470-480. (I.F. 6,6)
- Berko D., Tabachnick-Cherny S., Shental-Bechor D., Cascio P., Mioletti S., Levy Y., Admon A., Ziv T., Tirosh B., Goldberg A.L., Navon A. The direction of protein entry into the proteasome determines the variety of products and depends on the force needed to unfold its two termini. Molecular Cell, 2012, 48: 1-11. (I.F. 14,2)
- Cenci S., Oliva L., Cerruti F., Milan E., Bianchi G., Raule M., Mezghrani A., Pasqualetto E., Sitia R., Cascio P. Pivotal Advance: Protein synthesis modulates responsiveness of differentiating and malignant plasma cells to proteasome inhibitors. Journal of Leukocyte Biology, 2012, 92: 921-931. (I.F. 5)
- Favole A., Cascio P., Cerruti F., Sereno A., Tursi M., Tomatis A., Cristina Della Beffa C., Ferrone S., Bollo E. MHC Class I-releated antigen-processing machinery component defects in feline mammary carcinoma. Translational Oncology, 2012, 5: 48-55. (I.F. 3,4)
- Reina R., Juganaru M.M., Profiti M., Cascio P., Cerruti F., Bertolotti L., De Meneghi D., Amorena B., Rosati S. Immunological parameters in goats experimentaly infected with SRLV genotype E, strain Roccaverano. Veterinary Immunology and Immunopathology, 2011, 139: 237-244. (I.F. 2,2)
- Urru S., Veglianese P., De Luigi A., Fumagalli E., Erba E., Gonella Diaza R., Davoli E., Borsello T., Forloni G., Pengo N., Monzani E., Cascio P., Cenci S., Sitia R., Salmona M. A new fluorogenic peptide determines proteasome activity in single cells. Journal of Medicinal Chemistry, 2010, 53: 7452–7460. (I.F. 5,2)
- Cerruti F., Martano M., Morello E., Buracco P., Cascio P. Proteasomes are not a target for doxorubicin in feline injection-site sarcoma. Journal of Comparative Pathology, 2010, 143: 164-172. (I.F. 1,5)
- Bianchi G., Oliva L., Cascio P., Pengo N., Fontana F., Cerruti F., Orsi A., Pasqualetto E., Mezghrani A., Calbi V., Palladini G., Giuliani N., Anderson K. C., Sitia R., Cenci S. The proteasome load vs. capacity balance determines apoptotic sensitivity of multiple myeloma cells to proteasome inhibition. Blood, 2009, 113: 3040-49. (I.F. 10,6)
- Santoni de Sio F.R., Gritti A., Cascio P., Neri M., Sampaolesi M., Galli C., Luban J. and Naldini L. Lentiviral Vector Gene Transfer is Limited by the Proteasome at Post-Entry Steps in Various Types of Stem Cells. Stem Cells, 2008, 26: 2142-2152. (I.F. 7,7)
- Cascio P., Oliva L., Cerruti F., Mariani E., Pasqualetto E., Cenci S. and Sitia R. Dampening Ab responses using proteasome inhibitors following in vivo B cell activation. European Journal of Immunology, 2008, 38: 658-667. (I.F. 4,9)
- Cerruti F., Martano M., Petterino C., Bollo E., Morello E., Bruno R., Buracco P. and Cascio P. Enhanced expression of γ-interferon induced antigen processing machinery components in a spontaneously occurring cancer. Neoplasia, 2007, vol. 9, n.11: 960-969. (I.F. 5,7)
- Cenci S.*, Mezghrani A.*, Cascio P.*, Bianchi G., Cerruti F., Fra A., Lelouard H., Masciarelli S., Mattioli L., Oliva L., Orsi A., Pasqualetto E., Pierre P., Ruffato E., Tagliavacca L. and Sitia R. Progressively impaired proteasomal capacity during terminal plasma cell differentiation. The EMBO Journal, 2006, 25: 1104-13. (I.F. 10,1) (*Joint first authors)
- Santoni de Sio F.R., Cascio P., Zingale A., Gasparini M. and Naldini L. Proteasome activity restricts lentiviral gene transfer into hematopoietic stem cells and is down-regulated by cytokines that enhance transduction. Blood, 2006, 107: 4257-65. (I.F. 10,4)
- Petterino C., Martano M., Cascio P., Cerruti F., Martini M., Morello E., Bruno R., Castagnaro M., Buracco P. Immunohistochemical Study of STAT3 Expression in Feline Injection-site Fibrosarcomas. J. Comp. Pathology, 2006, 134: 91-100. (I.F. 0,9)
- Cascio P. and Goldberg A.L. Preparation of hybrid (19S-20S-PA28) proteasome complexes and analysis of peptides generated during protein degradation. Methods in Enzymology, 2005, 398: 336-352. (I.F. 1,7)
- Cheroni C., Peviani M., Cascio P., De Biasi S., Monti C. and Bendotti C. Accumulation of human SOD1 and ubiquitinated deposits in the spinal cord of SOD1G93A mice during motor neuron disease progression correlates with a decrease of proteasome. Neurobiology of Disease, 2005, 18: 509-22. (I.F. 4)
- Martano , Morello E., Ughetto M., Iussich S., Petterino C., Cascio P., Buracco P. Surgery alone versus surgery and doxorubicin for the treatment of feline injection-site sarcomas: a report on 69 cases. The Veterinary Journal, 2005, 170: 84-90. (I.F. 1,7)
- Goldberg A.L, Cascio P., Saric T., Rock K. The importance of the proteasome and subsequent proteolytic steps in the generation of antigenic peptides. Mol. Immunol., 2002, 39: 147-164. (I.F. 2,4)
- Cascio P., Call M., Petre BM., Walz T., Goldberg A.L. Properties of the hybrid form of the 26S proteasome containing both 19S and PA28 complexes. The EMBO Journal, 2002, 21: 2636-45. (I.F. 10,7)
- Cascio P., Hilton C., Kisselev A., Rock K., Goldberg A.L. 26S proteasomes and immunoproteasomes produce mainly N-extended versions of an antigenic peptide. The EMBO Journal, 2001, 20: 2357-2366. (I.F. 12,5)
- Kohler A., Cascio P., Leggete D.S., Woo K.M., Goldberg A.L., Finley D. The Axial Channel of the Proteasome Core Particle Is Gated by Rpt2 ATPase and Controls Both Substrate Entry and Product Release. Molecular Cell, 2001, 7: 1143-1152. (I.F. 16,6)
- Benaroudj N., Tarcsa E., Cascio P., Goldberg A.L. The unfolding of substrates and ubiquitin-independent protein degradation by proteasomes. Biochimie, 2001, 83: 311-318. (I.F. 2,7)
- Mo X.Y., Cascio P., Lemerise K., Goldberg A.L., Rock K. Distinct Proteolytic Processes Generate the C and N Termini of MHC Class I-binding Peptides. The Journal of Immunology, 1999, 163: 5851-5859. (I.F. 7,1)
- Piccinini M., Merighi A., Bruno R., Cascio P., Curto M., Mioletti S., Ceruti C., Rinaudo MT. Affinity purification and characterization of protein gene product 9.5 (PGP9.5) from retina. Biochemical Journal, 1996, 318: 711-6. (I.F. 3,6)
Insegnamenti
- Biochimica (VET0006)
Laurea Magistrale a Ciclo Unico in Medicina Veterinaria - Biochimica applicata (VET0006B)
Laurea Magistrale a Ciclo Unico in Medicina Veterinaria - Biochimica generale (VET0006A)
Laurea Magistrale a Ciclo Unico in Medicina Veterinaria - Elementi di propedeutica alla biochimica e biochimica (SVE0193)
Corso di Laurea in Tecniche di assistenza veterinaria - TAVET - Propedeutica alla biochimica e biochimica degli animali in allevamento (SVE0219)
Corso di Laurea in Sistemi zootecnici sostenibili - SIZOS
Temi di ricerca
Strictly controlled degradation of the proteome is a key factor in maintaining cellular homeostasis and allows a rapid and effective response to a variety of different stress challenges. The central element of the regulatory degradative network that integrates different stimuli and signals is the proteasome, a macromolecular machine designed to selectively remove specific proteins according to the variable needs of the cell. Not surprisingly, given the extreme complexity and interdependence of the pathways involved, the proteolytic system has evolved with a modular organization based on a central hydrolytic element, the 20S proteasome, to which alternative regulatory modules can associate to fine-tune its activity.
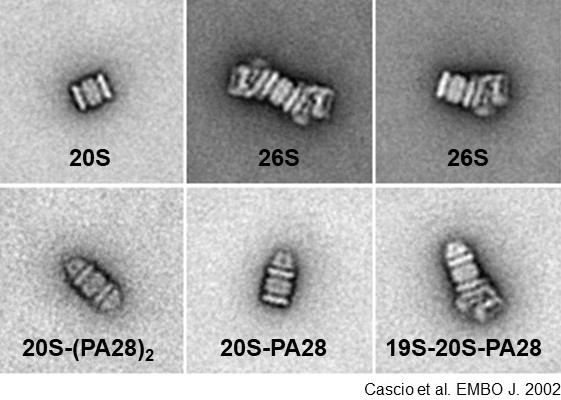
Average images of different proteasome forms obtained from electron micrographs of negatively stained particles
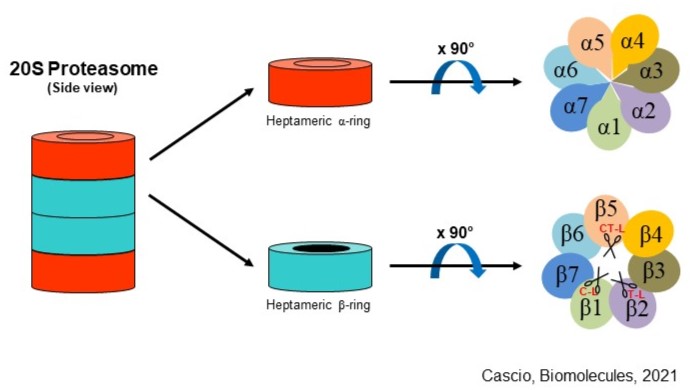
Schematic structure of 20S proteasome. Proteolytic active sites of β1, β2, and β5 subunits are depicted as scissors. C-L: caspase-like; T-L: trypsin-like; CT-L: chymotrypsin-like.
One of the complexes that activate the 20S proteasome is the 19S regulatory particle that, once associated with one or both ends of the 20S, leads to the formation of the 26S proteasome. The 19S regulatory cap performs the task of recognizing, unfolding, and translocating protein substrates within the protease via an ATP-dependent mechanism (i.e. energy-requiring).
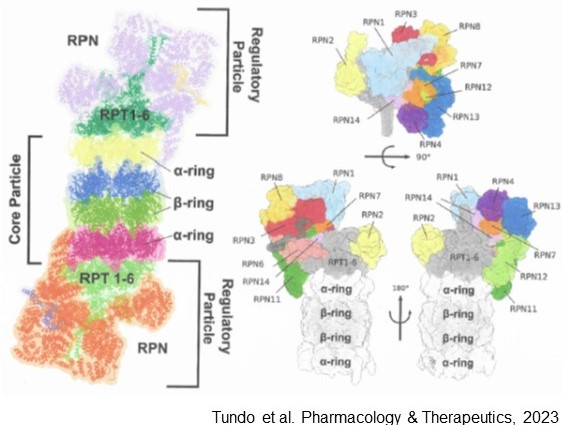
Overall organization of the proteasome 26S particle. Left: the “Core Particle” (20S proteasome) is represented as protein ribbons, in yellowand magenta the two α-subunit rings, in blue and green the two β-subunit rings. The two regulatory particles (19S), attached on both ends of the 20S particle, are represented as protein ribbons. The group of regulatory AAA-ATPases (Rpt1–6) are colored in dark and pale green, the non-ATPase regulators (Rpn) are colored in violet and orange. Right: close-up of the 26S regulatory particlewith the various non-ATPase subunits highlighted and labelled. The particle is shown fromvarious point of view: on the top panel it is shown fromthe top; on the lower panel two opposite side views are shown.
Another important family of 20S proteasome regulators is that of the ATP- and ubiquitin-independent PA28 activators (also called 11S, REG or PMSE), which in vertebrates is formed by the three highly homologous subunits α, β, and γ. PA28 monomers form ring-shaped ~200 kDa multimeric complexes that bind, in an ATP-independent manner, to the two ends of the 20S proteasome and profoundly modifies its peptidase, and in the case of PA28γ also its proteolytic, activities. In this regard, a recent paper from our lab demonstrated that diverse cellular functions of PA28γ (e.g. regulation of cell growth and proliferation, apoptosis, chromatin structure and organization, response to DNA damage) seem to depend on its ability to markedly enhance degradation rates of completely or largely unfolded proteins (IDPs, Intrinsically Disordered Proteins) by 20S proteasome.
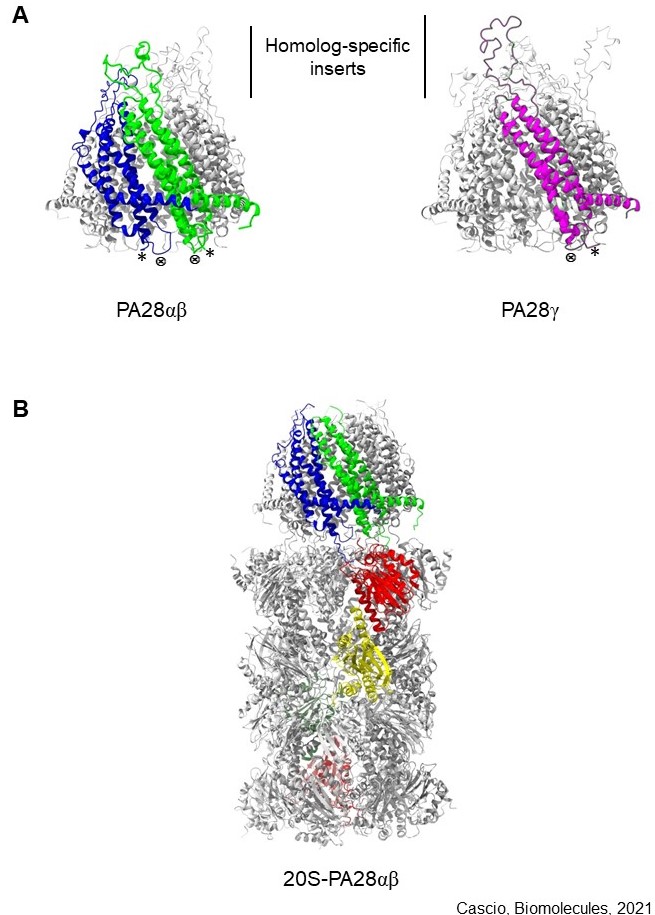
Model structure of PA28αβ , PA28γ and 20S-PA28αβ complexes. (A) Structure of the heteroheptameric PA28 αβ (left) and homoheptameric PA28γ (right). One (blue), one (green) and one (magenta) subunit are highlighted. * indicates C-terminus; ⊗ shows activation loop. Structures were generated by ChimeraX. (B) Structure of 20S-PA28αβ proteasome. One α (blue) and one β (green) subunit of PA28 and α 1 (red) and β1 (yellow) subunits of 20S are highlighted. Structures were generated by ChimeraX.
Not surprisingly, perturbation of this delicate degradative network is involved in the onset of various pathological conditions (e.g. cancer, neurodegenerative diseases, inflammatory and autoimmune disorders, infections) and aging. Consequently, in recent years strategies for modulating this proteolytic pathway (mainly through molecules that inhibit key enzymatic activities of this degradative system) have been shown to be extremely useful for both research and therapeutic purposes. Our laboratory is strongly committed to characterizing from a functional, molecular and structural point of view the different forms of proteasome (i.e. constitutive proteasome, proteasome containing the subunits induced by γ-INF, proteasomes associated with different ATP-dependent and independent activators) in order to unveil their biochemical properties and biological functions. To this end, we make use of biochemical, bio-molecular, mass spectrometry and cell culture techniques and approaches. In particular, our studies aim to characterize the activity of the proteasomal proteolytic system in the degradation of completely or largely unfolded (IDPs), misfolded, aggregated, and amyloidogenic proteins, and to understand its role in the onset and development of different human and domestic animal pathologies and biological processes, such as cancer, anti-tumor immunological responses, neurodegeneration, aging.
Gruppi di ricerca
Progetti di ricerca
Attività in agenda
Organi
Ricevimento studenti
Any day by appointment




